What is the name of the methodology?
Dual-color DNA-PAINT single-particle tracking for visualizing membrane protein interactions
OI Name
Kristina Ganzinger
What is the methodology good for?
Single-molecule microscopy is a powerful way to test whether membrane proteins bind each other or bind to soluble ligands. However, before we developed DNA-PAINT based single-particle tracking (DNA-PAINT-SPT), observation times in these experiments were often limited by fluorophore photobleaching. By implementing simultaneous live cell dual-color DNA-PAINT-SPT, we can now quantify protein dimerization of (1) proteins reconstituted in supported lipid bilayers, or (2) living cell membranes, and even extract 2D kinetics and 2D KD values.
What is/are the main advantages of this methodology over related technologies?
DNA-PAINT-SPT allows more precise measurements and is compatible with samples already expressing the protein(s) of interest labelled with common protein tags for imaging (e.g. GFP, SNAPtag, ALFA tag…). So far, the protein-of-interest has to be genetically labelled, limiting this approach mostly to cell lines. The methodology is particularly suitable for lowly-expressed (membrane) proteins with copy numbers of ~1000-5000 per cell.
What are the most important limitations of the methodology?
We can only study dimerization of recombinantly-expressed proteins (with at least one bound to a supported lipid bilayer) or membrane proteins in live cells that can be tagged from the extracellular side.
What type of samples are compatible with methodology?
Cancer cell lines | Primary cells in culture | Organoids | Primary tissue |
Yes | possibly | no | no |
What future develops to the methodology are you planning, in any?
We are working on an intracellular version, so that we can detect dimerization inside the cell. The proof-of-principle experiments look promising, so stay tuned and get in touch if you have an interesting test case!
If someone outside your lab wants to use the methodology, what is the best option?
A) What do you need to provide them to make it work (select one or more)?
Unless you have access to a total internal reflection fluorescence (TIRF) microscopy with single-molecule sensitivity, we will likely need to perform the experiments in our lab, in collaboration. In any case, it would be best to get in touch as setting up the experiments is somewhat challenging if you do it for the first time.
B) Is there any minimal expertise/equipment others need to work with the methodology?
If you want to perform the experiments yourself, you need to be trained in TIRF microscopy and automated data analysis (we would be happy to train you for the analysis and provide our scripts).
Name one or more people in your lab that are experienced with the methodology
Tom Aarts, Chi Nguyen, Edo Vreeker
Who originally developed the methodology?
We developed the method together with the groups of Petra Schwille and Ralf Jungmann at the MPI Biochemistry in Martinsried, Germany, and published it:
Christian Niederauer, Chikim Nguyen, Miles Wang-Henderson, Johannes Stein, Sebastian Strauss, Alexander Cumberworth, Florian Stehr, Ralf Jungmann, Petra Schwille & Kristina A. Ganzinger, Dual-color DNA-PAINT single-particle tracking enables extended studies of membrane protein interactions, Nature Communications, 2023, https://doi.org/10.1038/s41467-023-40065-8
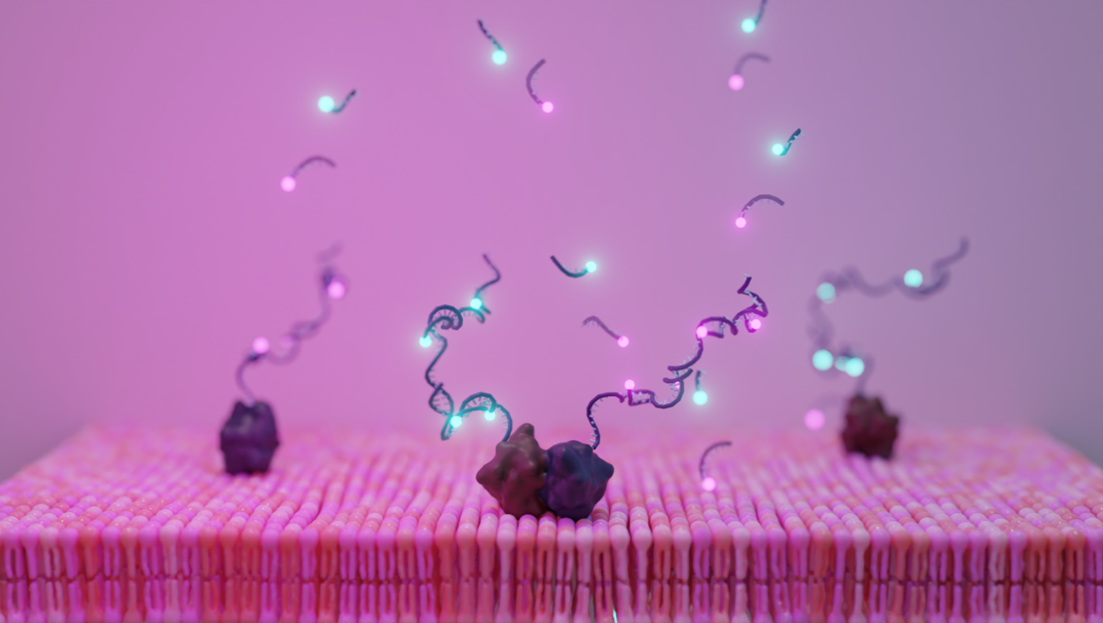
Self-renewing DNA-based fluorescent labels to visualize protein dimerization in single-molecule fluorescence experiments
Self-renewing DNA-based fluorescent labels to visualize protein dimerization in single-molecule fluorescence experiments

Simulations showing the detection (yellow spots) of co-diffusing, i.e. interacting, molecules
Simulations showing the detection (yellow spots) of co-diffusing, i.e. interacting, molecules